Welcome to the Hummon Research Group at the University of Notre Dame. Our research interests lie at the intersection of analytical chemistry and chemical biology, with a focus on cancer biology. Cancer is a complex disease, requiring sophisticated and systematic strategies to deliver knowledge that will lead to improved treatment options for patients. Individuals trained in a wide range of scientific topics and possessing a substantial breadth of knowledge are best positioned to tackle this challenge.
In the Hummon Research Group, we develop analytical methods to evaluate both the transcriptome and the proteome in cancer cells, while exploring the deregulation in cancer-associated signal transduction pathways.
We are part of the Department of Chemistry and Biochemistry at the University of Notre Dame and are affiliated with the Harper Cancer Research Institute. Our research lab is located in McCourtney Hall.
In the Hummon Research Group, we develop analytical methods to evaluate both the transcriptome and the proteome in cancer cells, while exploring the deregulation in cancer-associated signal transduction pathways.
We are part of the Department of Chemistry and Biochemistry at the University of Notre Dame and are affiliated with the Harper Cancer Research Institute. Our research lab is located in McCourtney Hall.
Current Projects:
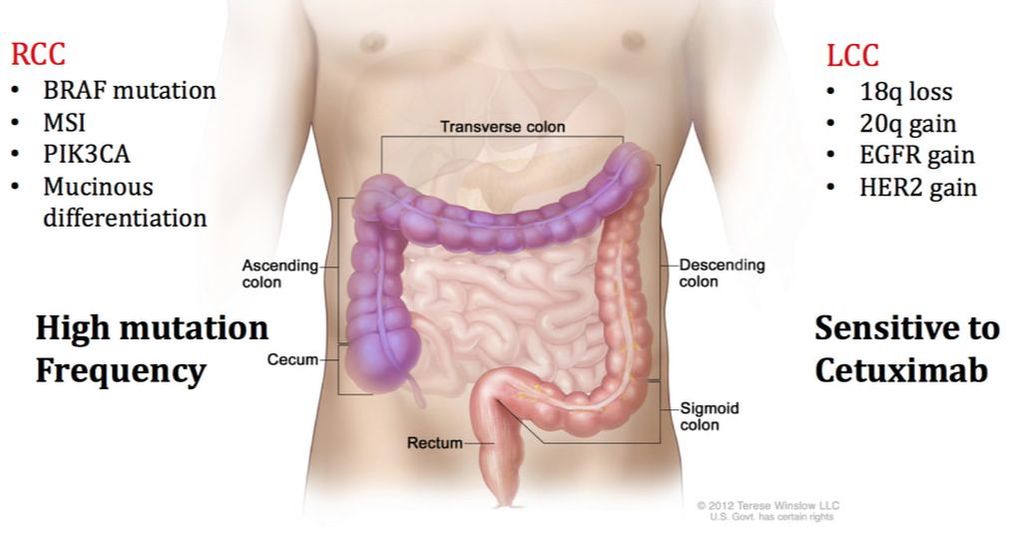
Proteomic Analysis of Right-Side and Left-Side Colon CancerNumerous studies have demonstrated that right-side colon cancer (RCC) and left-side colon cancer (LCC) exhibit distinct characteristics and should be treated as separate diseases. RCC spans the cecum, ascending colon, and transverse colon, while LCC includes the descending and sigmoid colon. RCC patients often have a poor prognosis compared to LCC patients because LCC patients respond more successfully to traditional FOLFOX and cetuximab treatments. RCC and LCC have distinct macroscopic and molecular characteristics. We are examining the proteomic differences found in RCC and LCC tumors to further understand the complexities of this disease.
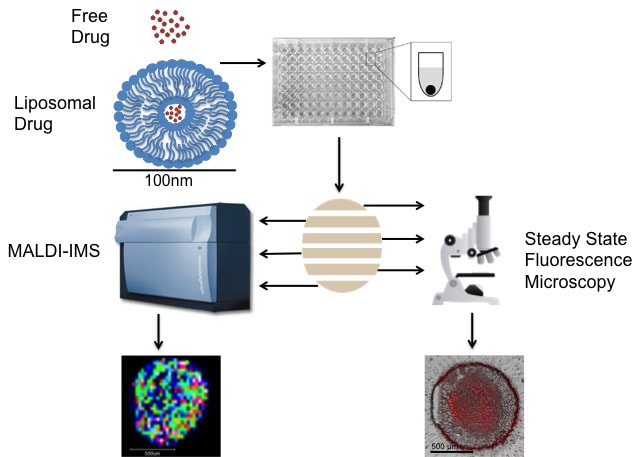
Analysis of Liposomal Drug Delivery SystemsCancer chemotherapeutics often fail to reach all diseased cells. To help solve this problem, researchers are investigating novel drug delivery systems. Liposomes are an attractive option due to their low toxicity, high biocompatibility, and potential to carry a large amount of a drug to the tumor site, all while avoiding being eliminated from the body. Using MALDI-IMS and fluorescence microscopy we analyze the spatial distribution of liposomal drugs in comparison to their free drug counterpart within spheroids.
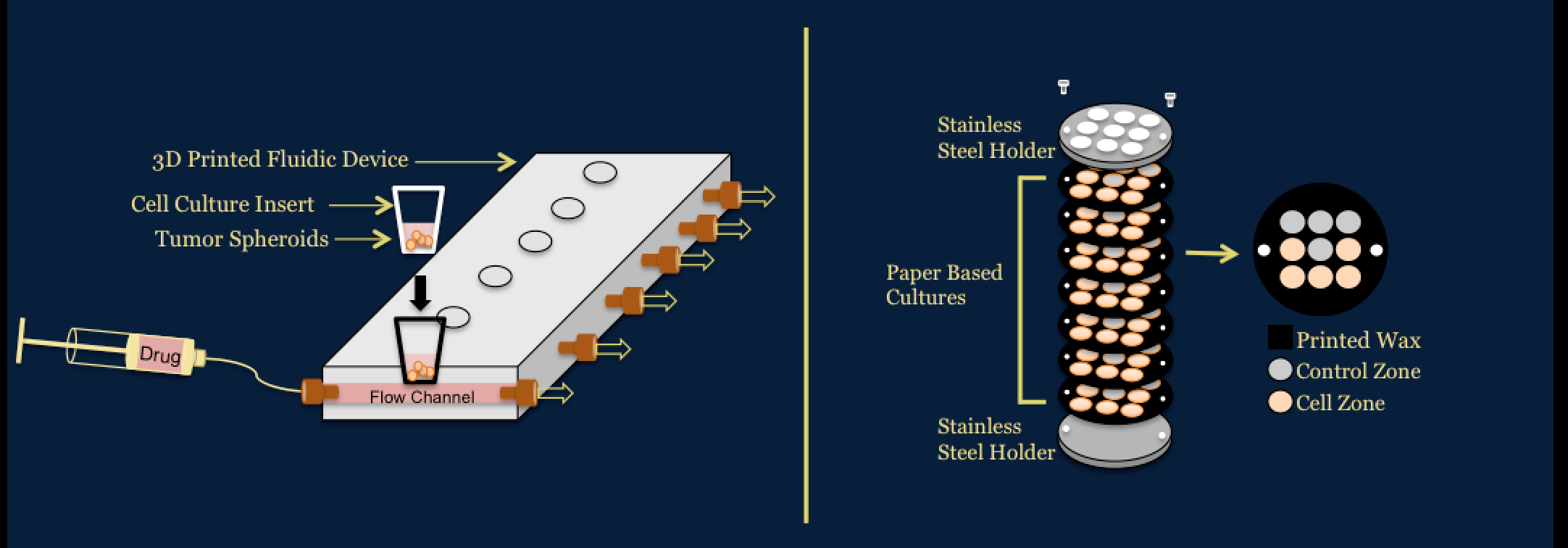
Investigation of Novel in vitro Platforms to Analyze the Efficacy and Toxicity of ChemotherapeuticsWe use a number of platforms to determine the efficacy and toxicity of chemotherapeutics for colorectal cancer. One platform utilizes a 3D printed fluidic device developed in the Dr. Dana Spence laboratory at Michigan State University to dynamically dose spheroids. The other platform utilizes paper based cultures developed in the Dr. Matthew Lockett laboratory at University of North Carolina at Chapel Hill. These systems allow for applications to preclinical trials area of drug development pipeline.
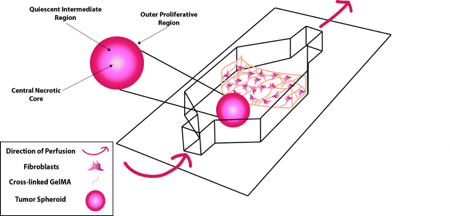
Modeling Metastasis and the Tumor MicroenvironmentThe combination of advanced 3D cell culture models and perfusion provides a powerful in vitro approach to modeling complex processes in disease. We are using this approach to investigate how migrating cancer cells interact with their surrounding environment.
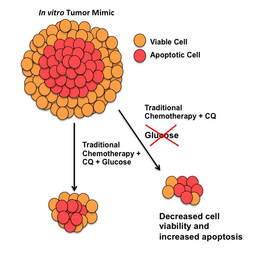
Investigating the molecular mechanisms behind how and why metastatic cells are able to escape the primary tumor mass and enter the blood stream is especially important. Advanced tumors shed hundreds of thousands to millions of cells during their lifetime. However, it has been shown that only about one out of every million cancer cells is successful at establishing metastatic disease. To investigate the discrepancy between the low success rate of metastatic cells and the high mortality rate of metastatic disease, we developed a fluidic device that combines a well characterized three dimensional tumor spheroid model system with multiple other cell types under perfusion. Confocal microscopy and imaging mass spectrometry (IMS) are used to examine the interactions between cell populations in order to learn how cancer cells respond to changes in the tumor microenvironment. This fully customizable, single channel device is made from poly(dimethylsiloxane) (PDMS) molded over a 3D-printed mold and is bonded to a glass slide using oxygen plasma treatment. Nutrient Restriction and Colorectal CancerThe use of short-term fasting is a potential cancer treatment that could be used in tandem with current cancer regimes to increase their potency. We set out to understand the molecular mechanisms behind nutrient restriction in CRC with the use quantitative proteomics. Quantitative proteomics is a tool to survey all of the proteins in a cell and their relative abundance between biological conditions. Using this technology, we are studying the changes that occur from the addition of glucose restriction and autophagy inhibition to treatment regimes for colorectal cancer.
|